CAPRI
From Wiki CEINGE
| Revision as of 16:52, 14 June 2007 (edit) Luca (Talk | contribs) m ← Previous diff |
Revision as of 20:24, 14 June 2007 (edit) (undo) Luca (Talk | contribs) Next diff → |
||
| Line 1: | Line 1: | ||
| CAPRI (Common Application Program Remote Interface) is a web interface for accessing all the major tools currently installed at CEINGE. It was developed at CEINGE [[The bioinformatics group|bionformatics lab]] and can be used only by registered users. | CAPRI (Common Application Program Remote Interface) is a web interface for accessing all the major tools currently installed at CEINGE. It was developed at CEINGE [[The bioinformatics group|bionformatics lab]] and can be used only by registered users. | ||
| - | CAPRI project was born to guarantee researchers an easy access to many widely used sequence analysis tools, such as [[BLAST]], [[FastA]] or even packages like [[EMBOSS]], which were originally developed on a command line interface. | + | CAPRI project was born to guarantee the researchers an easy access to many widely used sequence analysis tools, such as [[BLAST]], [[FastA]] or even packages like [[EMBOSS]], which were originally developed on a command line interface. |
| - | CAPRI resembles a typical local desktop application where the user selects the sequences and uses a number of menus to access the various functions but, in reality, | + | CAPRI resembles a typical local desktop application where the user selects the sequences and uses a number of menus to access the various functions but, in reality, it takes advantage of the processing power and the databases available on the CEINGE's remote servers. Unlike other similar tools CAPRI does not require to install any software. |
| Click [http://bioinfo.ceinge.unina.it/tools/winterf/capri/capri1.php here] to access to CAPRI. | Click [http://bioinfo.ceinge.unina.it/tools/winterf/capri/capri1.php here] to access to CAPRI. | ||
| Line 8: | Line 8: | ||
| A page of CAPRI is a web-page consisting of: | A page of CAPRI is a web-page consisting of: | ||
| - | * a | + | * a text-area (1), in which the user can insert and edit his data. Moreover, it is possible also to paste directly one or more sequences in this area. |
| - | * a lateral menu bar (2), which | + | * a lateral menu bar (2), which allows to: open other pages, load data from disk, servers or databases, copy the visible data to a new page and select that sequences that have to be displayed. |
| - | * a top menu bar (3), which | + | * a top menu bar (3), which allows to access the installed analysis tool and which can dynamically changes depending on the number of data. |
| - | Currently CAPRI implements five page | + | Currently CAPRI implements five kinds of page to analyze five different biological data: DNA, protein, alignment, Hidden Markov Model (HMM) and tree; each page contains links to access to all the programs available for the analysis of respective input data. |
| - | The result of | + | The result of each analysis is displayed in a new page, which type is in agreement with the output data: for example, DNA sequences are aligned within the 'DNA page' while the resulting alignment is provided in an 'alignment page'. |
| === DNA Page === | === DNA Page === | ||
| - | This is the CAPRI default page | + | This is the CAPRI default page that allows to analyze nucleic acid sequences like DNA or RNA. Top menu bar changes if one, two or more sequences are inserted and analyzed at the same time, indicating which tools are available for the analysis. |
| === Protein Page === | === Protein Page === | ||
| - | This page | + | This page allows the analysis of proteic sequences. Top menu bar changes if one, two or more sequences are inserted and analyzed at the same time, indicating which tools are available for the analysis. |
| === Alignment Page === | === Alignment Page === | ||
| - | + | With this page it is possible to analyze sequence alignments. | |
| === HMM Page === | === HMM Page === | ||
| This page allows the analysis of Hidden Markov Models (some information about HMM can be found [http://en.wikipedia.org/wiki/Hidden_Markov_model here]). | This page allows the analysis of Hidden Markov Models (some information about HMM can be found [http://en.wikipedia.org/wiki/Hidden_Markov_model here]). | ||
| === TREE Page === | === TREE Page === | ||
| This page allow the analysis of phylogenetic tree. | This page allow the analysis of phylogenetic tree. | ||
Revision as of 20:24, 14 June 2007
CAPRI (Common Application Program Remote Interface) is a web interface for accessing all the major tools currently installed at CEINGE. It was developed at CEINGE bionformatics lab and can be used only by registered users. CAPRI project was born to guarantee the researchers an easy access to many widely used sequence analysis tools, such as BLAST, FastA or even packages like EMBOSS, which were originally developed on a command line interface. CAPRI resembles a typical local desktop application where the user selects the sequences and uses a number of menus to access the various functions but, in reality, it takes advantage of the processing power and the databases available on the CEINGE's remote servers. Unlike other similar tools CAPRI does not require to install any software. Click here to access to CAPRI.
Contents |
Tool description
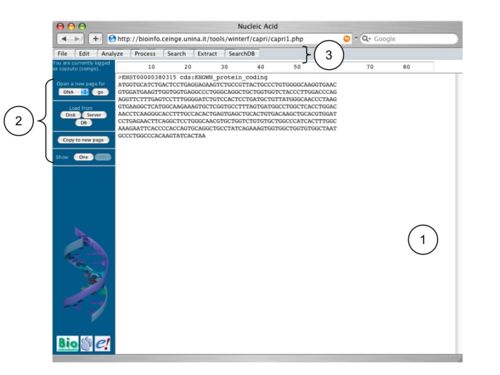
A page of CAPRI is a web-page consisting of:
- a text-area (1), in which the user can insert and edit his data. Moreover, it is possible also to paste directly one or more sequences in this area.
- a lateral menu bar (2), which allows to: open other pages, load data from disk, servers or databases, copy the visible data to a new page and select that sequences that have to be displayed.
- a top menu bar (3), which allows to access the installed analysis tool and which can dynamically changes depending on the number of data.
Currently CAPRI implements five kinds of page to analyze five different biological data: DNA, protein, alignment, Hidden Markov Model (HMM) and tree; each page contains links to access to all the programs available for the analysis of respective input data. The result of each analysis is displayed in a new page, which type is in agreement with the output data: for example, DNA sequences are aligned within the 'DNA page' while the resulting alignment is provided in an 'alignment page'.
DNA Page
This is the CAPRI default page that allows to analyze nucleic acid sequences like DNA or RNA. Top menu bar changes if one, two or more sequences are inserted and analyzed at the same time, indicating which tools are available for the analysis.
Protein Page
This page allows the analysis of proteic sequences. Top menu bar changes if one, two or more sequences are inserted and analyzed at the same time, indicating which tools are available for the analysis.
Alignment Page
With this page it is possible to analyze sequence alignments.
HMM Page
This page allows the analysis of Hidden Markov Models (some information about HMM can be found here).
TREE Page
This page allow the analysis of phylogenetic tree.