KINWEB
From Wiki CEINGE
| Revision as of 17:28, 20 June 2007 (edit) Mauro (Talk | contribs) ← Previous diff |
Revision as of 17:30, 20 June 2007 (edit) (undo) Mauro (Talk | contribs) (→Available annotations) Next diff → |
||
| Line 9: | Line 9: | ||
| Many computational analyses have been performed on the complete set of human kinome, including: | Many computational analyses have been performed on the complete set of human kinome, including: | ||
| *InterProScan analysis, in order to study domain distribution and combination in the various families; | *InterProScan analysis, in order to study domain distribution and combination in the various families; | ||
| - | *Comparative analysis of all gene regions and subsequent computational analysis, aimed at identifying CSTs, with particular focusing on those corresponding to | + | *Comparative analysis of all gene regions and subsequent computational analysis, aimed at identifying CSTs, with particular focusing on those corresponding to unannotated exons; |
| *Structural analysis, including the putative presence of transmembrane alpha helices, and the cystein propensity to participate into a disulfide bridge. | *Structural analysis, including the putative presence of transmembrane alpha helices, and the cystein propensity to participate into a disulfide bridge. | ||
| - | All the | + | All the analysis results have been stored into the KINWEB database and can be accessed from the web interface. |
Revision as of 17:30, 20 June 2007
Kinweb database is a collection of human protein kinases encoded in the human genome (human kinome) and relative annotations. Human kinase genes and their products have been extensively annotated: functional domains, secondary and tertiary structure of each gene product are reported, together with the full list of conserved sequence elements (CSTs), identified by comparative analysis of human kinase genes with their murine counterparts.
How to reach and use KINWEB
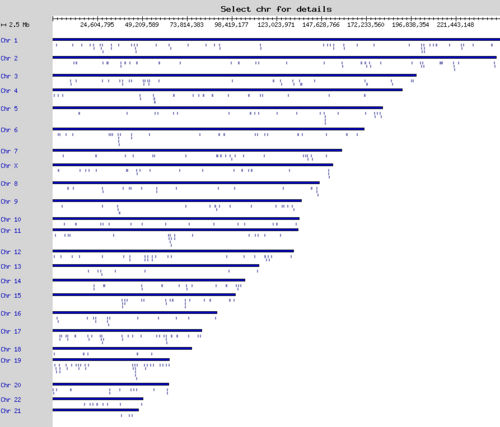
The database is available at http://kinweb.ceinge.unina.it: kinases can be searched by domain combinations and the relative genes may be viewed in a graphic browser at various level of magnification up to gene organization on the full chromosome set.
Available annotations
Many computational analyses have been performed on the complete set of human kinome, including:
- InterProScan analysis, in order to study domain distribution and combination in the various families;
- Comparative analysis of all gene regions and subsequent computational analysis, aimed at identifying CSTs, with particular focusing on those corresponding to unannotated exons;
- Structural analysis, including the putative presence of transmembrane alpha helices, and the cystein propensity to participate into a disulfide bridge.
All the analysis results have been stored into the KINWEB database and can be accessed from the web interface.